Lineweaver–Burk Plot
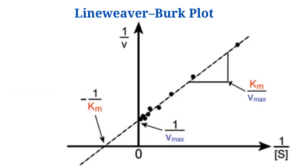
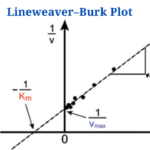
The Lineweaver–Burk plot (also known as the double reciprocal plot) is a graphical representation of enzyme kinetics used to determine the Michaelis-Menten parameters Vmax (maximum reaction velocity) and Km (Michaelis constant).
- Because Vmax is attained at endless substrate concentration, estimating Vmax and thus Km from a hyperbolic plot is impossible.
- Due to this issue, Lineweaver and Burk modified the Michaelis–Menten equation into a straight line equation.
- The Lineweaver–Burk plot (also known as the double reciprocal plot) is a pictorial depiction of the Lineweaver–Burk equation of enzyme kinetics, which was presented by Hans Lineweaver and Dean Burk in 1934.
- This figure is derived from the Michaelis–Menten equation and is denoted as follows:
where V represents the reaction velocity (reaction rate), Km denotes the Michaelis–Menten constant, Vmax denotes the maximum reaction velocity, and [S] denotes the substrate concentration.
- It produces a straight line with a y-axis intercept of 1/Vmax and an x-axis intercept of Km/Vmax. The slope of the line is equal to Km/Vmax.
- Vmax and Km can be calculated experimentally by measuring V0 at various substrate concentrations. Then, for 1/V0 vs 1/[S], a double reciprocal or Lineweaver–Burk plot is generated.
- A Lineweaver–Burk plot can be used to discriminate between competitive and noncompetitive reversible enzyme inhibitors.
- It is an effective method for determining how an inhibitor interacts to an enzyme.
- If V0 is measured at multiple substrate concentrations in the presence of a fixed dose of inhibitor, a Lineweaver–Burk plot can be used to detect competitive inhibition.
- On the Lineweaver–Burk plot, a competitive inhibitor increases the slope of the line and changes the intercept on the x-axis (because Km is increased), but leaves the intercept on the y-axis unaltered (since Vmax remains constant).
- Noncompetitive inhibition can also be identified on a Lineweaver–Burk plot because it raises the slope of the experimental line and changes the intercept on the y-axis (due to a drop in Vmax), but leaves the intercept on the x-axis unaltered (since Km remains constant).
Advantages:
-
Linearizes the Michaelis-Menten equation for easier analysis.
-
Allows clear determination of Vmax and Km.
Disadvantages:
-
Can amplify errors at low substrate concentrations (since reciprocal transformation exaggerates small values).
-
Other plots (e.g., Eadie-Hofstee) may be better for certain datasets.
Uses of Lineweaver–Burk Plot
-
Identifying enzyme inhibition types:
-
Competitive inhibition: Alters Km but not Vmax.
-
Non-competitive inhibition: Alters Vmax but not Km.
-
Uncompetitive inhibition: Alters both Km and Vmax.
-
Lineweaver-Burk Plot Worked Example
Experimental data for an enzyme-catalyzed reaction:
| [S] (mM) | v (μM/min) |
|---|---|
| 1.0 | 12.0 |
| 2.0 | 20.0 |
| 4.0 | 30.0 |
| 8.0 | 40.0 |
| 10.0 | 44.0 |
Step 1: Calculate 1/[S] and 1/v
| [S] (mM) | v (μM/min) | 1/[S] (mM⁻¹) | 1/v (min/μM) |
| 1.0 | 12.0 | 1.0 | 0.083 |
| 2.0 | 20.0 | 0.5 | 0.050 |
| 4.0 | 30.0 | 0.25 | 0.033 |
| 8.0 | 40.0 | 0.125 | 0.025 |
| 10.0 | 44.0 | 0.1 | 0.023 |
Step 2: Plot Interpretation
The Lineweaver-Burk plot shows a straight line where:
- Y-intercept = 1/Vmax
- Slope = Km/Vmax
- X-intercept = -1/Km
Step 3: Determine Vmax and Km
1. Find Vmax from Y-intercept:
Experimental y-intercept ≈ 0.02 min/μM
1/Vmax = 0.02 → Vmax = 1/0.02 = 50 μM/min
2. Find Km using slope:
Experimental slope ≈ 0.06 min
Slope = Km/Vmax → Km = Slope × Vmax = 0.06 × 50 = 3.0 mM
Alternative method using x-intercept:
Experimental x-intercept ≈ -0.33 mM⁻¹
-1/Km = -0.33 → Km = 1/0.33 ≈ 3.0 mM
Final Results
- Vmax = 50 μM/min
- Km = 3.0 mM
Conclusion
The enzyme has a maximum velocity of 50 μM/min and a Michaelis constant (Km) of 3.0 mM, meaning it reaches half-maximal velocity at 3 mM substrate concentration.
Lineweaver–Burk Plot Citations
- David Hames and Nigel Hooper (2005). Biochemistry. Third ed. Taylor & Francis Group: New York.
- Smith, C. M., Marks, A. D., Lieberman, M. A., Marks, D. B., & Marks, D. B. (2005). Marks’ basic medical biochemistry: A clinical approach. Philadelphia: Lippincott Williams & Wilkins.
- https://en.wikipedia.org/wiki/Lineweaver%E2%80%93Burk_plot
Related Posts
- Lineweaver–Burk Plot with Example
- Cell Wall: Definition, Diagram, Structure And Functions
- Phylum Porifera: Classification, Characteristics, Examples
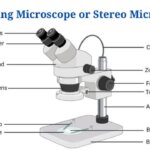
- Dissecting Microscope (Stereo Microscope) Definition, Principle, Uses, Parts
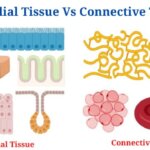
- Epithelial Tissue Vs Connective Tissue: Definition, 16+ Differences, Examples
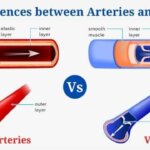
- 29+ Differences Between Arteries and Veins
- 31+ Differences Between DNA and RNA (DNA vs RNA)
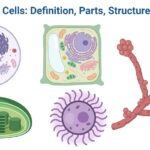
- Eukaryotic Cells: Definition, Parts, Structure, Examples
- Centrifugal Force: Definition, Principle, Formula, Examples
- Asexual Vs Sexual Reproduction: Overview, 18+ Differences, Examples
- Glandular Epithelium: Location, Structure, Functions, Examples
- 25+ Differences between Invertebrates and Vertebrates
- Cilia and Flagella: Definition, Structure, Functions and Diagram
- P-value: Definition, Formula, Table and Calculation
- Nucleosome Model of Chromosome